Search Results for author: Bowen Jing
Found 17 papers, 13 papers with code
Dirichlet Flow Matching with Applications to DNA Sequence Design
1 code implementation • 8 Feb 2024 • Hannes Stark, Bowen Jing, Chenyu Wang, Gabriele Corso, Bonnie Berger, Regina Barzilay, Tommi Jaakkola
Further, we provide distilled Dirichlet flow matching, which enables one-step sequence generation with minimal performance hits, resulting in $O(L)$ speedups compared to autoregressive models.
AlphaFold Meets Flow Matching for Generating Protein Ensembles
1 code implementation • 7 Feb 2024 • Bowen Jing, Bonnie Berger, Tommi Jaakkola
When trained and evaluated on the PDB, our method provides a superior combination of precision and diversity compared to AlphaFold with MSA subsampling.
Equivariant Scalar Fields for Molecular Docking with Fast Fourier Transforms
1 code implementation • 7 Dec 2023 • Bowen Jing, Tommi Jaakkola, Bonnie Berger
The runtime of our approach can be amortized at several levels of abstraction, and is particularly favorable for virtual screening settings with a common binding pocket.
Harmonic Self-Conditioned Flow Matching for Multi-Ligand Docking and Binding Site Design
1 code implementation • 9 Oct 2023 • Hannes Stärk, Bowen Jing, Regina Barzilay, Tommi Jaakkola
A significant amount of protein function requires binding small molecules, including enzymatic catalysis.
nnSAM: Plug-and-play Segment Anything Model Improves nnUNet Performance
1 code implementation • 29 Sep 2023 • Yunxiang Li, Bowen Jing, Zihan Li, Jing Wang, You Zhang
The recent developments of foundation models in computer vision, especially the Segment Anything Model (SAM), allow scalable and domain-agnostic image segmentation to serve as a general-purpose segmentation tool.
EigenFold: Generative Protein Structure Prediction with Diffusion Models
1 code implementation • 5 Apr 2023 • Bowen Jing, Ezra Erives, Peter Pao-Huang, Gabriele Corso, Bonnie Berger, Tommi Jaakkola
Protein structure prediction has reached revolutionary levels of accuracy on single structures, yet distributional modeling paradigms are needed to capture the conformational ensembles and flexibility that underlie biological function.
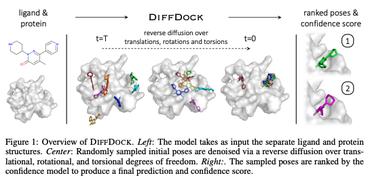
DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking
2 code implementations • 4 Oct 2022 • Gabriele Corso, Hannes Stärk, Bowen Jing, Regina Barzilay, Tommi Jaakkola
We instead frame molecular docking as a generative modeling problem and develop DiffDock, a diffusion generative model over the non-Euclidean manifold of ligand poses.
 Ranked #1 on
Blind Docking
on PDBbind
Ranked #1 on
Blind Docking
on PDBbind
Torsional Diffusion for Molecular Conformer Generation
1 code implementation • 1 Jun 2022 • Bowen Jing, Gabriele Corso, Jeffrey Chang, Regina Barzilay, Tommi Jaakkola
Molecular conformer generation is a fundamental task in computational chemistry.
Subspace Diffusion Generative Models
1 code implementation • 3 May 2022 • Bowen Jing, Gabriele Corso, Renato Berlinghieri, Tommi Jaakkola
Score-based models generate samples by mapping noise to data (and vice versa) via a high-dimensional diffusion process.
 Ranked #19 on
Image Generation
on CIFAR-10
Ranked #19 on
Image Generation
on CIFAR-10
Equivariant Graph Neural Networks for 3D Macromolecular Structure
2 code implementations • 7 Jun 2021 • Bowen Jing, Stephan Eismann, Pratham N. Soni, Ron O. Dror
Representing and reasoning about 3D structures of macromolecules is emerging as a distinct challenge in machine learning.
ATOM3D: Tasks On Molecules in Three Dimensions
3 code implementations • 7 Dec 2020 • Raphael J. L. Townshend, Martin Vögele, Patricia Suriana, Alexander Derry, Alexander Powers, Yianni Laloudakis, Sidhika Balachandar, Bowen Jing, Brandon Anderson, Stephan Eismann, Risi Kondor, Russ B. Altman, Ron O. Dror
We implement several classes of three-dimensional molecular learning methods for each of these tasks and show that they consistently improve performance relative to methods based on one- and two-dimensional representations.
Protein model quality assessment using rotation-equivariant, hierarchical neural networks
no code implementations • 27 Nov 2020 • Stephan Eismann, Patricia Suriana, Bowen Jing, Raphael J. L. Townshend, Ron O. Dror
Proteins are miniature machines whose function depends on their three-dimensional (3D) structure.
Learning from Protein Structure with Geometric Vector Perceptrons
3 code implementations • ICLR 2021 • Bowen Jing, Stephan Eismann, Patricia Suriana, Raphael J. L. Townshend, Ron Dror
Learning on 3D structures of large biomolecules is emerging as a distinct area in machine learning, but there has yet to emerge a unifying network architecture that simultaneously leverages the graph-structured and geometric aspects of the problem domain.
Rotation-Invariant Gait Identification with Quaternion Convolutional Neural Networks
1 code implementation • 4 Aug 2020 • Bowen Jing, Vinay Prabhu, Angela Gu, John Whaley
A desireable property of accelerometric gait-based identification systems is robustness to new device orientations presented by users during testing but unseen during the training phase.
Hierarchical, rotation-equivariant neural networks to select structural models of protein complexes
no code implementations • 5 Jun 2020 • Stephan Eismann, Raphael J. L. Townshend, Nathaniel Thomas, Milind Jagota, Bowen Jing, Ron O. Dror
Predicting the structure of multi-protein complexes is a grand challenge in biochemistry, with major implications for basic science and drug discovery.
SGVAE: Sequential Graph Variational Autoencoder
no code implementations • 17 Dec 2019 • Bowen Jing, Ethan A. Chi, Jillian Tang
Generative models of graphs are well-known, but many existing models are limited in scalability and expressivity.
Modeling Sensorimotor Coordination as Multi-Agent Reinforcement Learning with Differentiable Communication
no code implementations • 12 Sep 2019 • Bowen Jing, William Yin
Multi-agent reinforcement learning has shown promise on a variety of cooperative tasks as a consequence of recent developments in differentiable inter-agent communication.
 Multi-agent Reinforcement Learning
Multi-agent Reinforcement Learning
 reinforcement-learning
+1
reinforcement-learning
+1