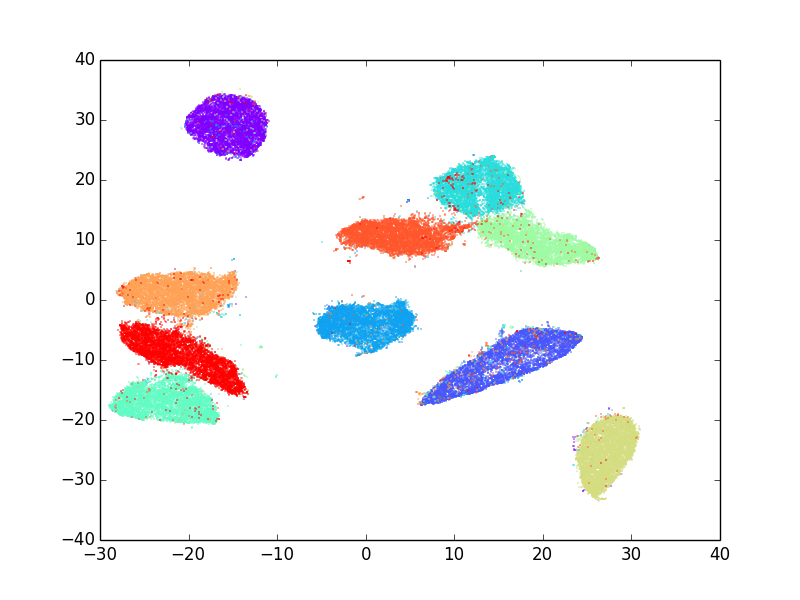
Visualizing Large-scale and High-dimensional Data
We study the problem of visualizing large-scale and high-dimensional data in a low-dimensional (typically 2D or 3D) space. Much success has been reported recently by techniques that first compute a similarity structure of the data points and then project them into a low-dimensional space with the structure preserved. These two steps suffer from considerable computational costs, preventing the state-of-the-art methods such as the t-SNE from scaling to large-scale and high-dimensional data (e.g., millions of data points and hundreds of dimensions). We propose the LargeVis, a technique that first constructs an accurately approximated K-nearest neighbor graph from the data and then layouts the graph in the low-dimensional space. Comparing to t-SNE, LargeVis significantly reduces the computational cost of the graph construction step and employs a principled probabilistic model for the visualization step, the objective of which can be effectively optimized through asynchronous stochastic gradient descent with a linear time complexity. The whole procedure thus easily scales to millions of high-dimensional data points. Experimental results on real-world data sets demonstrate that the LargeVis outperforms the state-of-the-art methods in both efficiency and effectiveness. The hyper-parameters of LargeVis are also much more stable over different data sets.
PDF Abstract