Protein-Ligand Complex Generator & Drug Screening via Tiered Tensor Transform
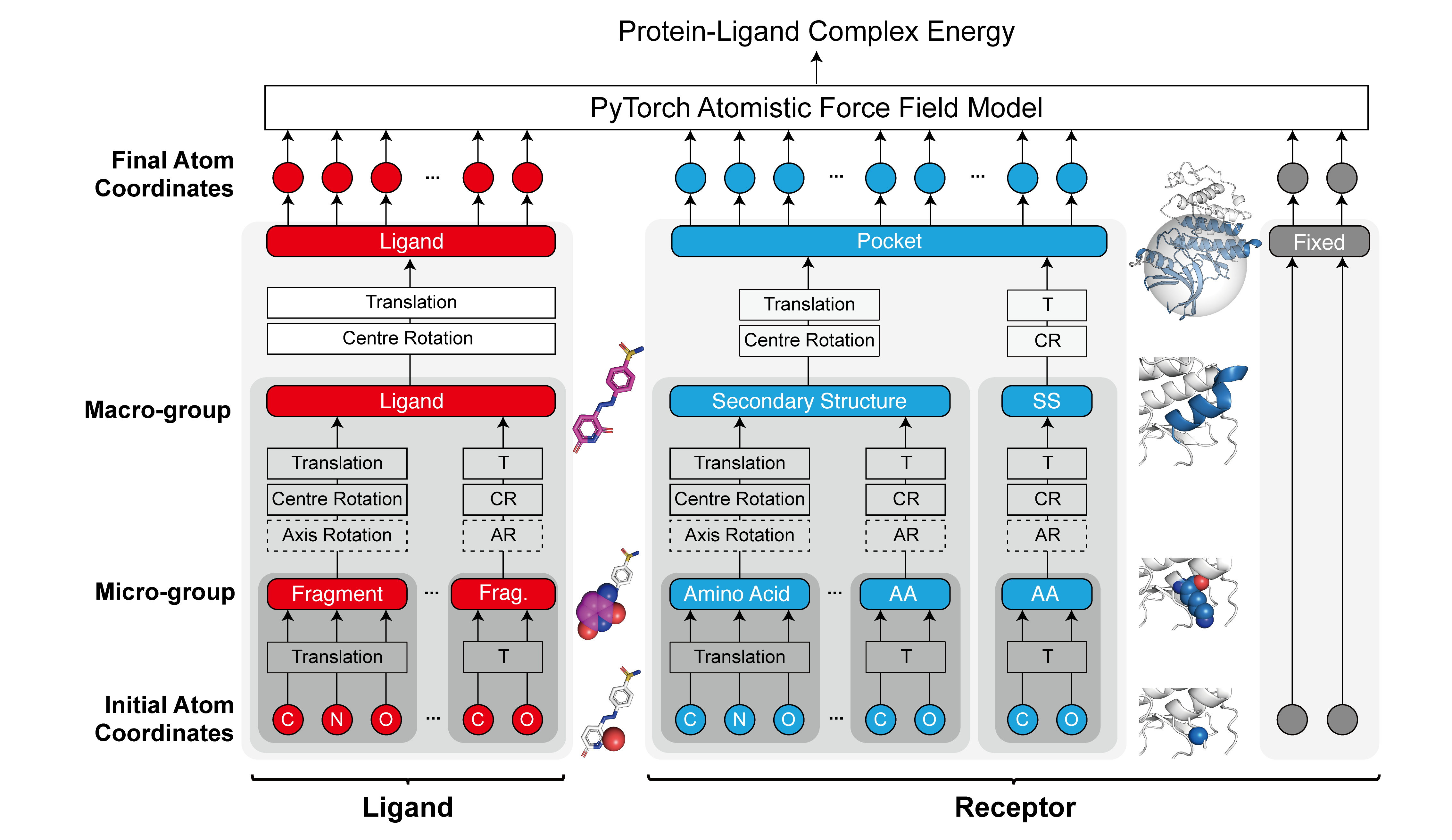
The generation of small molecule candidate (ligand) binding poses in its target protein pocket is important for computer-aided drug discovery. Typical rigid-body docking methods ignore the pocket flexibility of protein, while the more accurate pose generation using molecular dynamics is hindered by slow protein dynamics. We develop a tiered tensor transform (3T) algorithm to rapidly generate diverse protein-ligand complex conformations for both pose and affinity estimation in drug screening, requiring neither machine learning training nor lengthy dynamics computation, while maintaining both coarse-grain-like coordinated protein dynamics and atomistic-level details of the complex pocket. The 3T conformation structures we generate achieve significantly higher accuracy in active ligand classification than traditional ensemble docking using hundreds of experimental protein conformations. Furthermore, we demonstrate that 3T can be used to explore distant protein-ligand binding poses within the protein pocket. 3T structure transformation is decoupled from the system physics, making future usage in other computational scientific domains possible.
PDF Abstract