Learning a low dimensional manifold of real cancer tissue with PathologyGAN
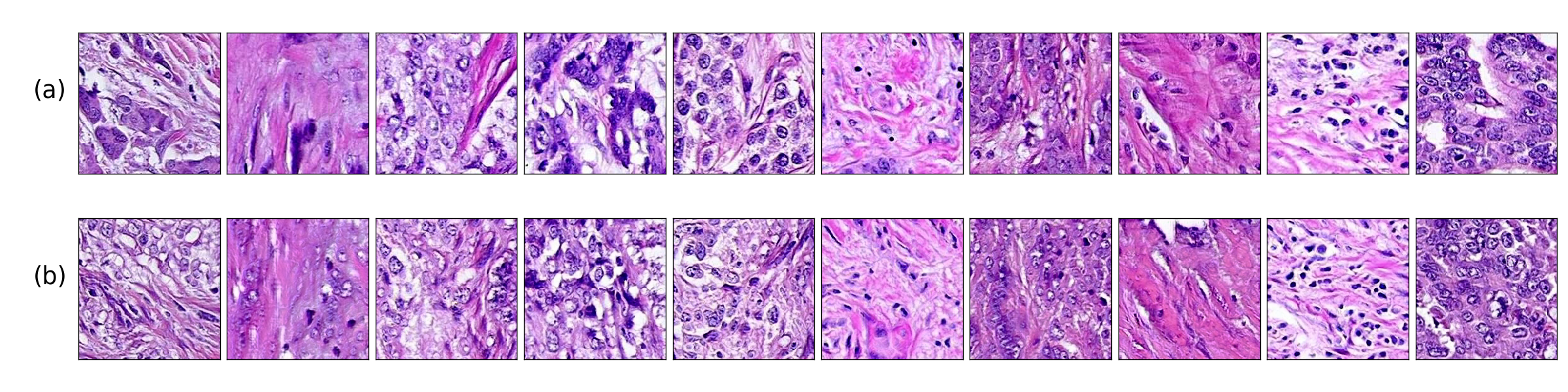
Application of deep learning in digital pathology shows promise on improving disease diagnosis and understanding. We present a deep generative model that learns to simulate high-fidelity cancer tissue images while mapping the real images onto an interpretable low dimensional latent space. The key to the model is an encoder trained by a previously developed generative adversarial network, PathologyGAN. We study the latent space using 249K images from two breast cancer cohorts. We find that the latent space encodes morphological characteristics of tissues (e.g. patterns of cancer, lymphocytes, and stromal cells). In addition, the latent space reveals distinctly enriched clusters of tissue architectures in the high-risk patient group.
PDF Abstract