FastSAM3D: An Efficient Segment Anything Model for 3D Volumetric Medical Images
Segment anything models (SAMs) are gaining attention for their zero-shot generalization capability in segmenting objects of unseen classes and in unseen domains when properly prompted. Interactivity is a key strength of SAMs, allowing users to iteratively provide prompts that specify objects of interest to refine outputs. However, to realize the interactive use of SAMs for 3D medical imaging tasks, rapid inference times are necessary. High memory requirements and long processing delays remain constraints that hinder the adoption of SAMs for this purpose. Specifically, while 2D SAMs applied to 3D volumes contend with repetitive computation to process all slices independently, 3D SAMs suffer from an exponential increase in model parameters and FLOPS. To address these challenges, we present FastSAM3D which accelerates SAM inference to 8 milliseconds per 128*128*128 3D volumetric image on an NVIDIA A100 GPU. This speedup is accomplished through 1) a novel layer-wise progressive distillation scheme that enables knowledge transfer from a complex 12-layer ViT-B to a lightweight 6-layer ViT-Tiny variant encoder without training from scratch; and 2) a novel 3D sparse flash attention to replace vanilla attention operators, substantially reducing memory needs and improving parallelization. Experiments on three diverse datasets reveal that FastSAM3D achieves a remarkable speedup of 527.38x compared to 2D SAMs and 8.75x compared to 3D SAMs on the same volumes without significant performance decline. Thus, FastSAM3D opens the door for low-cost truly interactive SAM-based 3D medical imaging segmentation with commonly used GPU hardware. Code is available at https://github.com/arcadelab/FastSAM3D.
PDF Abstract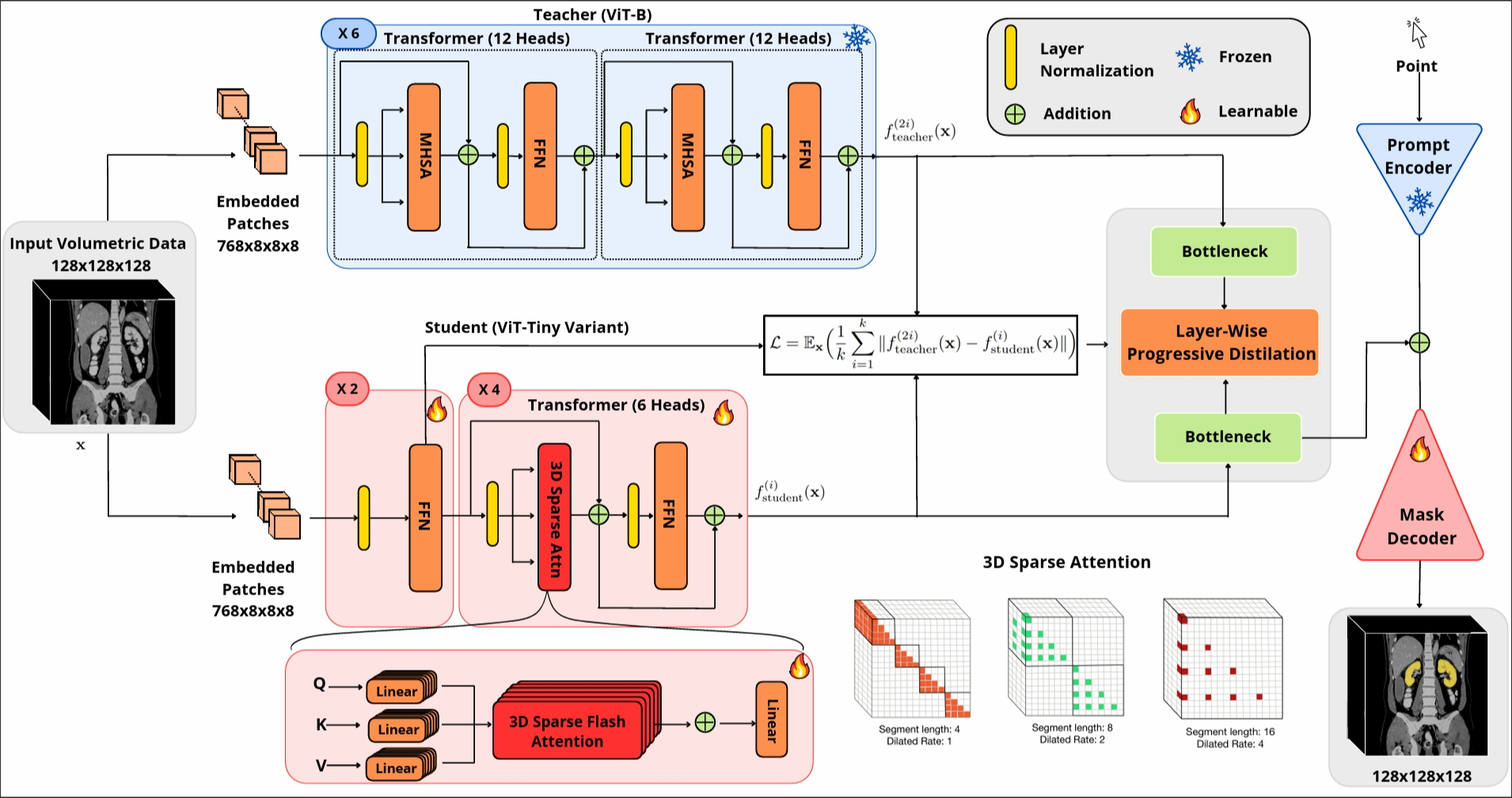





 AMOS
AMOS