3D-TransUNet for Brain Metastases Segmentation in the BraTS2023 Challenge
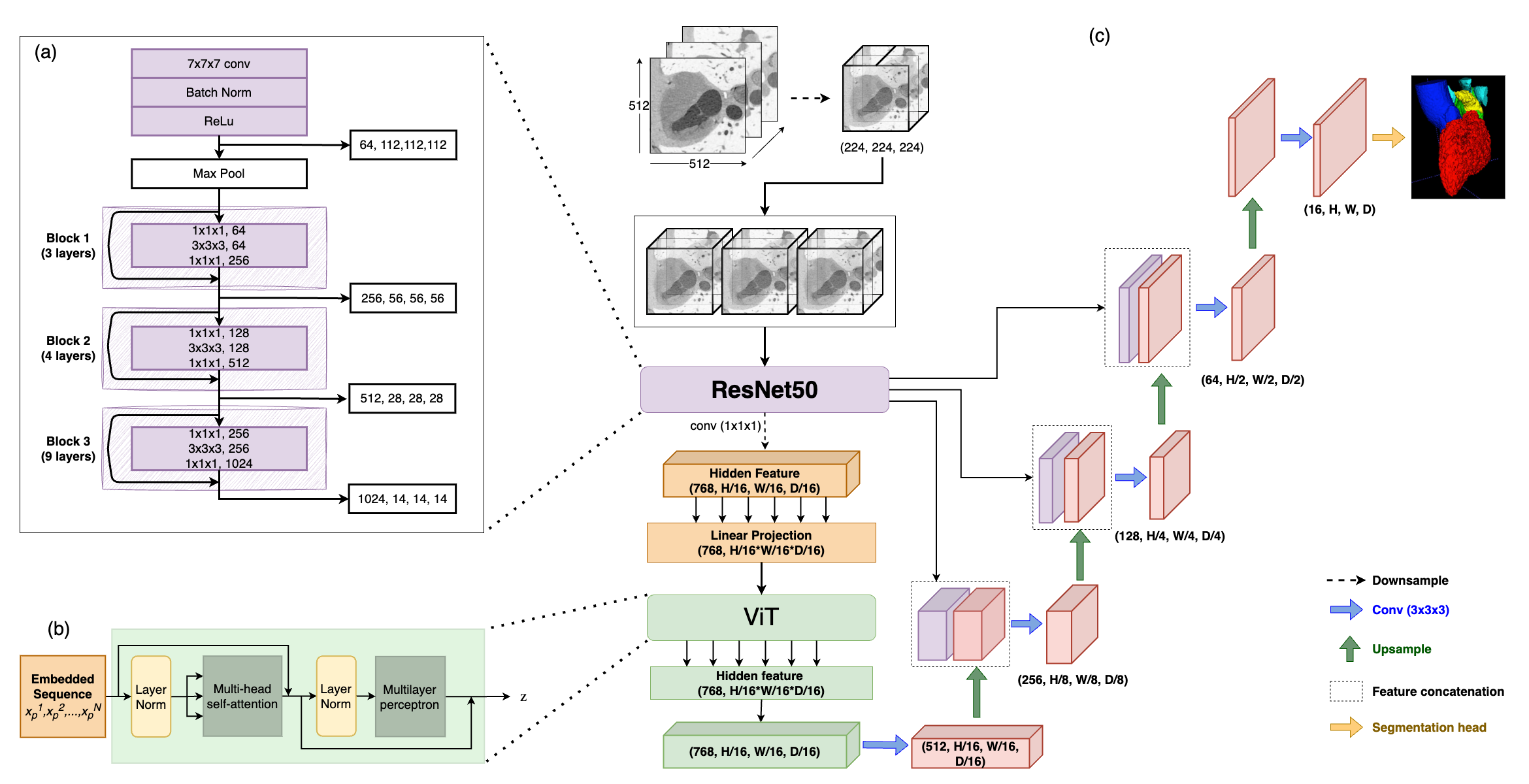
Segmenting brain tumors is complex due to their diverse appearances and scales. Brain metastases, the most common type of brain tumor, are a frequent complication of cancer. Therefore, an effective segmentation model for brain metastases must adeptly capture local intricacies to delineate small tumor regions while also integrating global context to understand broader scan features. The TransUNet model, which combines Transformer self-attention with U-Net's localized information, emerges as a promising solution for this task. In this report, we address brain metastases segmentation by training the 3D-TransUNet model on the Brain Tumor Segmentation (BraTS-METS) 2023 challenge dataset. Specifically, we explored two architectural configurations: the Encoder-only 3D-TransUNet, employing Transformers solely in the encoder, and the Decoder-only 3D-TransUNet, utilizing Transformers exclusively in the decoder. For Encoder-only 3D-TransUNet, we note that Masked-Autoencoder pre-training is required for a better initialization of the Transformer Encoder and thus accelerates the training process. We identify that the Decoder-only 3D-TransUNet model should offer enhanced efficacy in the segmentation of brain metastases, as indicated by our 5-fold cross-validation on the training set. However, our use of the Encoder-only 3D-TransUNet model already yield notable results, with an average lesion-wise Dice score of 59.8\% on the test set, securing second place in the BraTS-METS 2023 challenge.
PDF Abstract